HistoHelper
HistoHelper: WSI Inference & Predictive Morphology
1. Tl;dr
HistoHelper is an active research prototype exploring predictive morphology in oncology. The objective is to evaluate whether a Convolutional Neural Network can identify morphological signatures within primary skin lesions to predict specific organ tropism (e.g., pulmonary vs. cerebral metastasis).
- Frontend Interface: FastAPI, Uvicorn, Docker, Google Cloud Run
- Machine Learning Pipeline: PyTorch (MPS Hardware Accelerated), OpenCV, OpenSlide
- Model Architecture: ResNet18 (Feature Extraction) transitioning to Multiple Instance Learning.
- WSI Tiling Engine: Dynamic OpenCV Otsu’s Thresholding for glass filtration and spatial patching.
2. Phase 1 Validation: Patch-Level Feature Extraction
The V1 inference engine successfully classifies high-magnification histological patches. Utilising a custom HistoGradCAM interpreter hooked into ResNet18’s terminal convolutional layer, the model successfully generates localised heatmaps (e.g., highlighting dense dermal collagen bundles in benign samples), confirming it relies on biologically relevant cellular structures rather than background noise.
3. Phase 2: Whole Slide Image (WSI) Inference Pipeline
To bridge the gap between individual patches and clinical WSI diagnostics, a full ingestion and reconstruction pipeline was engineered:
- Dynamic Segmentation (
mvp_tiler.py): UtilisesOpenSlideto read gigapixel.svsbiopsies without memory exhaustion.OpenCVapplies Otsu’s thresholding to a low-resolution thumbnail to generate a binary tissue mask, successfully discarding 100% of blank glass and extracting only valid tissue coordinates. - Macroscopic Heatmap Reconstruction (
wsi_inference.py): The PyTorchDataLoaderpushes thousands of extracted patches through the frozen ResNet18 model in highly concurrent batches. The resulting probabilities are spatially mapped back to their original coordinates, generating a macroscopic topographical heatmap using an OpenCVCOLORMAP_JEToverlay.
4. Phase 2 Quantitative Validation (The Patch-Level Classifier)
To evaluate clinical viability, the V2 production weights (histohelper_production.pth) were evaluated against a holdout set of 2,000 unseen histology patches.
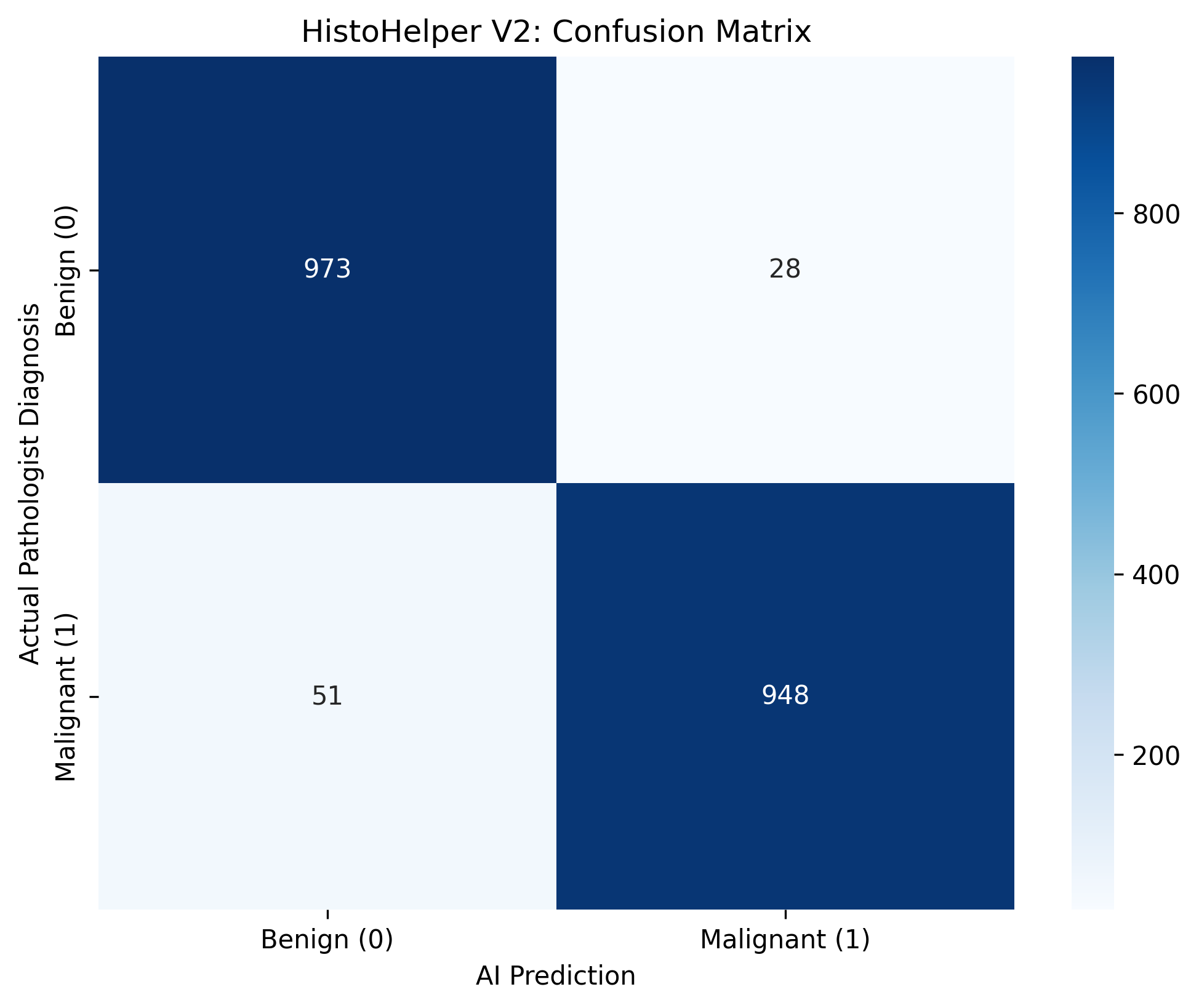
The model achieved an overall accuracy of 96.05%, demonstrating robust feature extraction capabilities:
- Sensitivity (Recall): 94.89% (948 True Positives vs 51 False Negatives).
- Specificity: 97.20% (973 True Negatives vs 28 False Positives).
5. Clinical Limitations & The Pivot to MIL
(See Commit 2f9df41 for the first WSI Heatmap Output)
While the extraction and reconstruction geometry is flawless, the resulting heatmap exposes the inherent limitations of patch-level classifiers on macroscopic WSIs:
- Domain Shift & Color Variance: The model suffers from “static” hallucination when evaluating out-of-distribution WSIs due to variations in H&E staining protocols and scanner lighting artifacts.
- The Context Deficit: Tumors grow in contiguous sheets, but a patch-level classifier evaluates a 224x224 microscopic keyhole in isolation, often flagging benign stroma as malignant due to localised noise.
The V3 Roadmap: To solve Patient Leakage and contextual fragmentation, the architecture is pivoting to Multiple Instance Learning. Instead of classifying individual patches, the WSI will be treated as a “bag” of instances. An Attention-Based MIL aggregator will be trained to assign weights to the most suspicious patches, allowing the network to evaluate the slide holistically before rendering a diagnostic prediction.